- Imprimer
- Partager
- Partager sur Facebook
- Partager sur LinkedIn
Séminaire / Équipe CoMHet
Le 19 janvier 2024
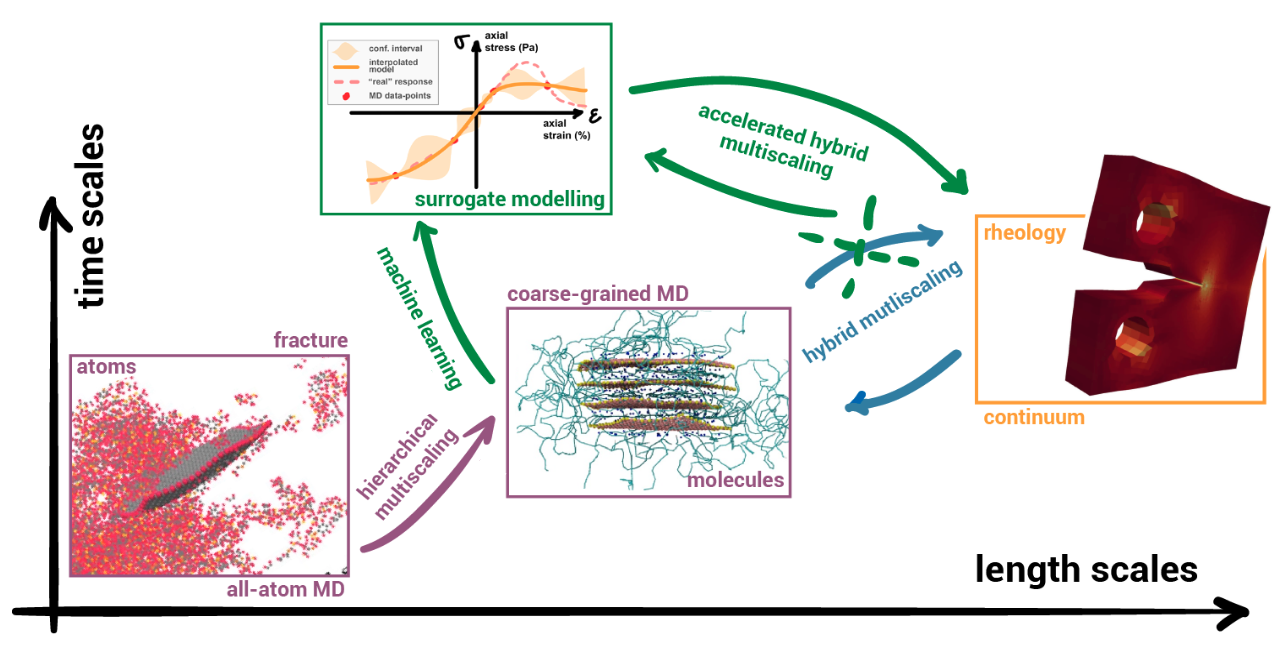
Probing mechanical properties of (bio)polymers with chemical specificity using molecular and multiscale simulation
Molecular dynamics simulations are ubiquitous in materials science, from drug discovery to design of advanced structural nanocomposites. Owing to the high-resolution of these simulations taking place at the atomic scale, predictions give access to data often complementary to experiments; be it characterisation of the nanostructure or even sophisticated instrumentation. I will begin the talk introducing the fundamentals of setting up molecular models and dynamics simulations to investigate the mechanics of materials. I will illustrate these aspects focusing on the collagen, the structural protein of choice in the human body and more largely the animal kingdom. I will present our recent investigations on the influence of hydration on the assembly of collagen microfibrils and the complex water dynamics within, trying to draw conclusions on the mechanical properties of the biopolymer.
While being quite popular, molecular dynamics simulations have several limitations (force field precision, ergodicity). Among these, the spectrum of spatiotemporal scales integrated within a simulation is extremely limited. This is particularly problematic when mechanical properties are of interest, as these emerge from the combination of scales ranging from the nanoscale (chemistry) to the macroscale (processing, testing). Unlike real-life experiments, all scales cannot be resolved simultaneously using computer simulations. I will give an overview of the existing multiscale simulation strategies: from rather cheap hierarchical to expensive concurrent approaches. I will illustrate their applicability with examples from my past research on the fracture of concrete under seismic loading and the dynamic behaviour of impacted shells of epoxy-graphene nanocomposites.
Date
10h30
Localisation
Galilée room 015

- Imprimer
- Partager
- Partager sur Facebook
- Partager sur LinkedIn
